Browse through our Journals...
OPTIMISATION OF BENZOFURAN / BENZOTHIOPHENE BIPHENYL AS PTPase 1B INHIBITOR USING QSAR MODEL
Darpan Kaushika, S.G. Kaskhedikarb & S.A. Khana
aDept. of Pharm. Chem., F/O Pharmacy, Jamia Hamdard, New Delhi-62.
bDept. of Pharmacy, S.G.S.I.T.S., Indore.
Insulin resistance can result from a defect in the insulin receptor signaling system at a site post binding of insulin to its receptor. Insulin resistance is associated with a defect in protein tyrosine phosphorylation in the insulin signal transduction cascade. PTPase enzyme dephosphorylates the active form of insulin receptor and thus attenuates its tyrosine kinase activity.
Insulin resistance is thus one of the obstacles which we confront while undergoing therapy for diabetes mellitus. A number of PTPase inhibitors has been designed and studied to overcome this problem, to gain insight into the structural and molecular requirements influencing the PTPase-1B inhibition activity, we here in describe the QSAR analysis of a set of structurally different compounds of PTPase inhibitors, for which it is conceivable to make assumption that they interact with the enzyme.
Experiment and discussion
Malamas et al reported seven series of compounds based on benzofuran/benzothiophene biphenyl moiety. We had performed the QSAR analysis of series III & IV which was having 7 & 10 compounds, out of which only 6 & 7 compounds in respective series could be subjected to 2-D QSAR analysis because of the non-availability of exact value of IC50 for some substituted compounds. 2D QSAR study was carried out in the following steps:
1). Calculations of physico-chemical constants from literature.
The values for the physicochemical constants for various substituents were determined from the literature. The determined parameters for series III include, Hansch constant (p), Molar Refractivity (h), Sigma/Hammet constant (s), Field effect (f) and Indicator value (I).
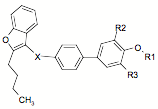
The substituent in the structure I:
For series III: IR1, IR2, IR3, πX, σX, ηX, fX
IR1 (Indicator parameter of R1 substituent)
IR1= 1, where -CH2COOH present at R1 position, otherwise 0.
IR2 (Indicator parameter for R2 substituent)
IR2 = 1, where H is at R2 position, otherwise, other substituent considered as 0.
IR3 (Indicator parameter for R3 substituent)
IR3 = 1, where H is at R3 position, otherwise, other substituent considered as 0.
πX, σX, ηX, fX are the physicochemical parameter for substitution X.
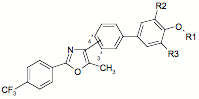
The substituents in structure II are as follows:
For series IV: IR1, πR1, σR1, ηR1, fR1, IR2, IR3 and Ia
IR1 (Indicator parameter for R1 substituent)
IR1=1, where -CH2COOH present at R1 position, otherwise 0.
πR1, σR1, ηR1, fR1 are the physicochemical parameter for IR1.
IR2 (Indicator parameter for R2 substituent)
IR2 = 1, where H is at R2 position, otherwise, other substituent considered as 0.
IR3 (Indicator parameter for R3 substituent)
IR3 = 1, where H present at R3 position, otherwise considered as 0.
Ip (Indicator parameter for POA)
Ip = 1, when POA is 4, rest other attachment considered as 0.
2) Determination of the correlation matrix:
The correlation matrix for series III & IV was determined separately using the program ‘SYSTAT’ (version 7.0). The most significant parameters for PTPase inhibiting activity were chosen on the basis of their correlation ship & Interco relationship.
3) Multiple regression analysis:
It was performed by using the program ‘SYSTAT’ for PTPase inhibiting activity, i.e. –logIC50 considered as dependent variable and IR1, IR2, IR3, πX, σX, ηX, fX & IR1,πR1, σR1, ηR1, fR1, IR2, IR3 and Ia were selected as the independent variables separately for series III & IV.
The significant equation obtained for series III & IV are equation 1 & 2 respectively:
-logIC50 = 0.595(± 0.297) πX – 0.594(± 0.271) IR2 + 0.212(± 0.225)
n = 6, r = 0.82, s = 0.223, F = 3.088 ………. (1)
-logIC50 = 0.511(± 0.197) IR3 - 0.467(± 0.149)
n = 7, r = 0.758, s = 0.258, F = 6.743 ………. (2)
The data showed more than the critical values of the Pearson Product-Moment Correlation coefficient for both the eq. (1) & (2) along with overall significance level better than 95 % than the tabulated value of F in eq. (2) but the first one doesn’t comply this. So on this basis it can’t be possible to conclude from eq. (1) since equation found to be non significant, so we just laid down our stress on structure II that any substitution at R3 position is going to decrease the activity of lead moiety for the PTPase 1B activity because of possible steric hindrance of bulky group with the aceto moiety attached to the biphenyl ring.
Finally, it can be concluded that the work presented here will play an important role in understanding the relationship of physicochemical parameters with structure and biological activity of the PTPase 1B inhibitor and will help in choosing the suitable substituent for getting the active compound with maximum potency.
Bibliography:
1) Haring, HU. (1991). Diabetologia, 34, 848-861.
2) Byon, JCH, Kusari, AB. and Kusari, J. (1998). Mol.Cell.Biochem., 182, 101-108.
3) Malamas, MS, Sredy, J, Moxham, C, Katz, A, W.Xu, McDevitt, R, Adebayo, FO, Sawicki, DR, Seestaller, L, Sullivan, D and Taylor; JR.(2000) J. Med. Chem. 43, 1293-1310.
4) Hansch, C and Leo, A. (1979) Substituent constants for correlation analysis in chemistry and biology, john wiley and sons, New York, 48.
5) Systat, (version 7.0). S.P.S.S., Inc., 944, North Michigan Avenue, Chicago, IL 60611.U.S.
Click
on these links to visit our Journals:
Psychiatry
On-Line
Dentistry On-Line | Vet
On-Line | Chest Medicine
On-Line
GP
On-Line | Pharmacy
On-Line | Anaesthesia
On-Line | Medicine
On-Line
Family Medical
Practice On-Line
Home • Journals • Search • Rules for Authors • Submit a Paper • Sponsor us
All pages in this site copyright ©Priory Lodge Education Ltd 1994-